Section 5.3 — Bayesian difference between means#
This notebook contains the code examples from Section 5.3 Bayesian difference between means from the No Bullshit Guide to Statistics.
See also t-test.ipynb
Notebook setup#
# load Python modules
import os
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# Figures setup
plt.clf() # needed otherwise `sns.set_theme` doesn"t work
from plot_helpers import RCPARAMS
RCPARAMS.update({"figure.figsize": (5, 3)}) # good for screen
# RCPARAMS.update({"figure.figsize": (5, 1.6)}) # good for print
sns.set_theme(
context="paper",
style="whitegrid",
palette="colorblind",
rc=RCPARAMS,
)
# High-resolution please
%config InlineBackend.figure_format = "retina"
# Where to store figures
DESTDIR = "figures/bayesian/dmeans"
<Figure size 640x480 with 0 Axes>
# set random seed for repeatability
np.random.seed(42)
#######################################################
Example movies genres#
see https://www.andrewheiss.com/blog/2019/01/29/diff-means-half-dozen-ways/
see also https://bookdown.org/content/3686/metric-predicted-variable-on-one-or-two-groups.html#two-groups
movies = pd.read_csv("../datasets/movies.csv")
movies.head()
| title | year | rating | genre | genre_numeric | |
|---|---|---|---|---|---|
| 0 | Blowing Wild | 1953 | 5.6 | Action | 1 |
| 1 | No Way Back | 1995 | 5.2 | Action | 1 |
| 2 | New Jack City | 1991 | 6.1 | Action | 1 |
| 3 | Noigwon | 1983 | 4.2 | Action | 1 |
| 4 | Tarzan and the Jungle Boy | 1968 | 5.2 | Action | 1 |
movies.groupby("genre")["rating"].mean()
genre
Action 5.2845
Comedy 5.9670
Name: rating, dtype: float64
from scipy.stats import ttest_ind
actions = movies[movies["genre"]=="Action"]["rating"]
comedies = movies[movies["genre"]=="Comedy"]["rating"]
ttest_ind(actions, comedies, equal_var=True)
TtestResult(statistic=-4.47525173500199, pvalue=9.976981171112132e-06, df=398.0)
ttest_ind(actions, comedies, equal_var=False)
TtestResult(statistic=-4.47525173500199, pvalue=9.978285839671782e-06, df=397.7995256063933)
import bambi as bmb
WARNING (pytensor.tensor.blas): Using NumPy C-API based implementation for BLAS functions.
# Model formula
levels = ["Comedy", "Action"]
formula = bmb.Formula("rating ~ 1 + C(genre, levels=levels)")
# Choose custom priors
priors = {
"Intercept": bmb.Prior("Normal", mu=0, sigma=5),
"C(genre, levels=levels)": bmb.Prior("Normal", mu=0, sigma=1)
}
# Build model
model_eq = bmb.Model(formula, priors=priors, data=movies)
# Get model description
print(model_eq)
Formula: rating ~ 1 + C(genre, levels=levels)
Family: gaussian
Link: mu = identity
Observations: 400
Priors:
target = mu
Common-level effects
Intercept ~ Normal(mu: 0.0, sigma: 5.0)
C(genre, levels=levels) ~ Normal(mu: 0.0, sigma: 1.0)
Auxiliary parameters
sigma ~ HalfStudentT(nu: 4.0, sigma: 1.559)
# model_eq.build()
# model_eq.graph()
# Fit the model using 1000 on each chain
idata_eq = model_eq.fit(draws=1000)
Auto-assigning NUTS sampler...
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [sigma, Intercept, C(genre, levels=levels)]
Sampling 2 chains for 1_000 tune and 1_000 draw iterations (2_000 + 2_000 draws total) took 2 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
import arviz as az
az.summary(idata_eq, stat_focus="median")
| median | mad | eti_3% | eti_97% | mcse_median | ess_median | ess_tail | r_hat | |
|---|---|---|---|---|---|---|---|---|
| sigma | 1.527 | 0.035 | 1.431 | 1.630 | 0.002 | 2550.767 | 1789.0 | 1.00 |
| Intercept | 5.955 | 0.072 | 5.756 | 6.165 | 0.002 | 2413.684 | 1619.0 | 1.00 |
| C(genre, levels=levels)[Action] | -0.665 | 0.103 | -0.950 | -0.376 | 0.004 | 2405.330 | 1230.0 | 1.01 |
Regression, assuming unequal variances#
levels = ["Comedy", "Action"]
formula_uneq = bmb.Formula("rating ~ 1 + C(genre,levels=levels)", "sigma ~ C(genre,levels=levels)")
priors = {
"Intercept": bmb.Prior("Normal", mu=0, sigma=5),
"C(genre, levels=levels)": bmb.Prior("Normal", mu=0, sigma=1),
"sigma": {"C(genre, levels=levels)": bmb.Prior("Cauchy", alpha=0, beta=1)},
}
# Build model
model_uneq = bmb.Model(formula_uneq, priors=priors, data=movies)
# Get model description
print(model_uneq)
# model_uneq.build()
# model_uneq.graph()
Formula: rating ~ 1 + C(genre,levels=levels)
sigma ~ C(genre,levels=levels)
Family: gaussian
Link: mu = identity
sigma = log
Observations: 400
Priors:
target = mu
Common-level effects
Intercept ~ Normal(mu: 0.0, sigma: 5.0)
C(genre, levels=levels) ~ Normal(mu: 0.0, sigma: 1.0)
target = sigma
Common-level effects
sigma_Intercept ~ Normal(mu: 0.0, sigma: 1.0)
sigma_C(genre, levels=levels) ~ Cauchy(alpha: 0.0, beta: 1.0)
idata_uneq = model_uneq.fit(draws=1000)
Auto-assigning NUTS sampler...
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [Intercept, C(genre, levels=levels), sigma_Intercept, sigma_C(genre, levels=levels)]
Sampling 2 chains for 1_000 tune and 1_000 draw iterations (2_000 + 2_000 draws total) took 3 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
az.summary(idata_uneq, stat_focus="median")
| median | mad | eti_3% | eti_97% | mcse_median | ess_median | ess_tail | r_hat | |
|---|---|---|---|---|---|---|---|---|
| Intercept | 5.960 | 0.076 | 5.752 | 6.166 | 0.004 | 3498.346 | 1634.0 | 1.0 |
| C(genre, levels=levels)[Action] | -0.669 | 0.103 | -0.962 | -0.377 | 0.003 | 3252.640 | 1633.0 | 1.0 |
| sigma_Intercept | 0.432 | 0.033 | 0.339 | 0.528 | 0.001 | 3047.433 | 1401.0 | 1.0 |
| sigma_C(genre, levels=levels)[Action] | -0.021 | 0.051 | -0.151 | 0.122 | 0.001 | 3341.266 | 1478.0 | 1.0 |
np.exp(az.summary(idata_uneq, stat_focus="median").loc["sigma_C(genre, levels=levels)[Action]","median"])
0.9792189645694596
BEST#
levels = ["Comedy", "Action"]
formula_best = bmb.Formula("rating ~ 1 + C(genre,levels=levels)", "sigma ~ C(genre,levels=levels)")
priors = {
"Intercept": bmb.Prior("Normal", mu=0, sigma=5),
"C(genre, levels=levels)": bmb.Prior("Normal", mu=0, sigma=1),
"sigma": {"C(genre, levels=levels)": bmb.Prior("Cauchy", alpha=0, beta=1)},
"nu": bmb.Prior("Exponential", lam=1/29),
}
# Build model
model_best = bmb.Model(formula_best, priors=priors, family="t", data=movies)
# Get model description
print(model_best)
# model_best.build()
# model_best.graph()
Formula: rating ~ 1 + C(genre,levels=levels)
sigma ~ C(genre,levels=levels)
Family: t
Link: mu = identity
sigma = log
Observations: 400
Priors:
target = mu
Common-level effects
Intercept ~ Normal(mu: 0.0, sigma: 5.0)
C(genre, levels=levels) ~ Normal(mu: 0.0, sigma: 1.0)
Auxiliary parameters
nu ~ Exponential(lam: 0.0345)
target = sigma
Common-level effects
sigma_Intercept ~ Normal(mu: 0.0, sigma: 1.0)
sigma_C(genre, levels=levels) ~ Cauchy(alpha: 0.0, beta: 1.0)
idata_best = model_best.fit(draws=1000)
Auto-assigning NUTS sampler...
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [nu, Intercept, C(genre, levels=levels), sigma_Intercept, sigma_C(genre, levels=levels)]
Sampling 2 chains for 1_000 tune and 1_000 draw iterations (2_000 + 2_000 draws total) took 4 seconds.
There were 3 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
az.summary(idata_best, stat_focus="median")
| median | mad | eti_3% | eti_97% | mcse_median | ess_median | ess_tail | r_hat | |
|---|---|---|---|---|---|---|---|---|
| nu | 32.494 | 13.937 | 10.378 | 108.382 | 0.602 | 1576.809 | 1333.0 | 1.0 |
| Intercept | 5.980 | 0.076 | 5.779 | 6.178 | 0.003 | 1918.068 | 1332.0 | 1.0 |
| C(genre, levels=levels)[Action] | -0.685 | 0.103 | -0.984 | -0.391 | 0.006 | 1843.197 | 1325.0 | 1.0 |
| sigma_Intercept | 0.390 | 0.039 | 0.271 | 0.496 | 0.002 | 1400.381 | 1359.0 | 1.0 |
| sigma_C(genre, levels=levels)[Action] | -0.003 | 0.051 | -0.143 | 0.149 | 0.002 | 2045.051 | 1521.0 | 1.0 |
BEST with priors on variables instead of difference#
levels = ["Comedy", "Action"]
formula_best2 = bmb.Formula("rating ~ 0 + C(genre,levels=levels)",
"sigma ~ 0 + C(genre,levels=levels)")
priors = {
"C(genre, levels=levels)": bmb.Prior("TruncatedNormal", mu=6, sigma=2, lower=1, upper=10),
"sigma": {"C(genre, levels=levels)": bmb.Prior("Cauchy", alpha=0, beta=1)},
"nu": bmb.Prior("Exponential", lam=1/29),
}
# Build model
model_best2 = bmb.Model(formula_best2, priors=priors, family="t", data=movies)
# Get model description
print(model_best2)
# model_best2.build()
# model_best2.graph()
Formula: rating ~ 0 + C(genre,levels=levels)
sigma ~ 0 + C(genre,levels=levels)
Family: t
Link: mu = identity
sigma = log
Observations: 400
Priors:
target = mu
Common-level effects
C(genre, levels=levels) ~ TruncatedNormal(mu: 6.0, sigma: 2.0, lower: 1.0, upper: 10.0)
Auxiliary parameters
nu ~ Exponential(lam: 0.0345)
target = sigma
Common-level effects
sigma_C(genre, levels=levels) ~ Cauchy(alpha: 0.0, beta: 1.0)
idata_best2 = model_best2.fit(draws=1000)
Auto-assigning NUTS sampler...
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [nu, C(genre, levels=levels), sigma_C(genre, levels=levels)]
Sampling 2 chains for 1_000 tune and 1_000 draw iterations (2_000 + 2_000 draws total) took 3 seconds.
There were 5 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
az.summary(idata_best2, stat_focus="median")
| median | mad | eti_3% | eti_97% | mcse_median | ess_median | ess_tail | r_hat | |
|---|---|---|---|---|---|---|---|---|
| nu | 30.528 | 13.632 | 9.989 | 107.766 | 0.676 | 1830.640 | 1200.0 | 1.0 |
| C(genre, levels=levels)[Comedy] | 5.985 | 0.064 | 5.785 | 6.188 | 0.002 | 2167.875 | 1407.0 | 1.0 |
| C(genre, levels=levels)[Action] | 5.293 | 0.075 | 5.094 | 5.504 | 0.003 | 1731.895 | 1327.0 | 1.0 |
| sigma_C(genre, levels=levels)[Comedy] | 0.385 | 0.040 | 0.261 | 0.491 | 0.001 | 1808.968 | 1068.0 | 1.0 |
| sigma_C(genre, levels=levels)[Action] | 0.383 | 0.037 | 0.278 | 0.488 | 0.002 | 1854.532 | 1214.0 | 1.0 |
np.exp(az.summary(idata_best2, stat_focus="median").loc["sigma_C(genre, levels=levels)[Action]","median"])
1.4666780300683986
Example from original BEST paper#
Data taken from BESTexample-original.R in BEST.zip
via https://web.archive.org/web/20170708173718/https://www.indiana.edu/~kruschke/BEST/
Steps following Matti Vuorre’s blog post see also src notebook.
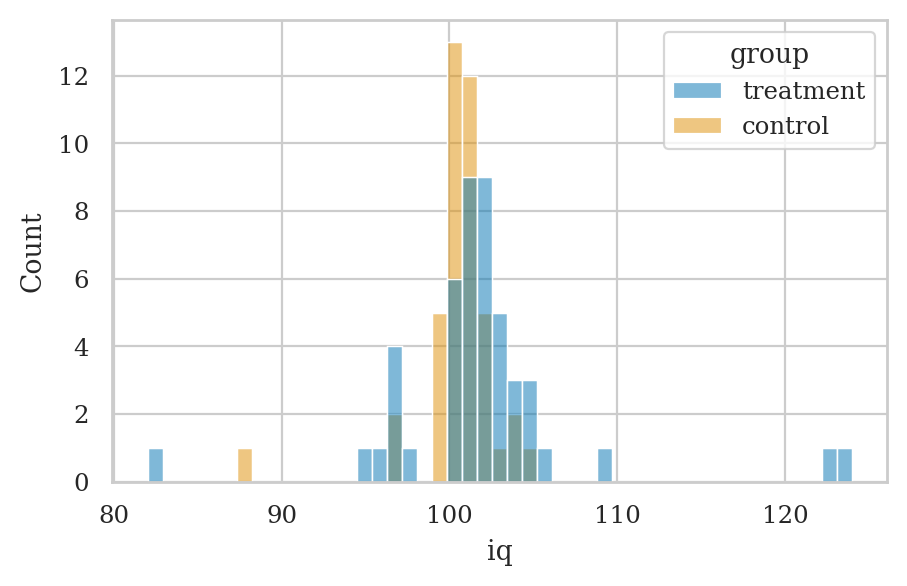
Data#
treated = np.array([101, 100, 102, 104, 102, 97, 105, 105, 98, 101, 100, 123, 105,
103, 100, 95, 102, 106, 109, 102, 82, 102, 100, 102, 102, 101,
102, 102, 103, 103, 97, 97, 103, 101, 97, 104, 96, 103, 124,
101, 101, 100, 101, 101, 104, 100, 101])
controls = np.array([ 99, 101, 100, 101, 102, 100, 97, 101, 104, 101, 102, 102, 100,
105, 88, 101, 100, 104, 100, 100, 100, 101, 102, 103, 97, 101,
101, 100, 101, 99, 101, 100, 100, 101, 100, 99, 101, 100, 102,
99, 100, 99])
d = pd.DataFrame({
"group": ["treatment"]*len(treated) + ["control"]*len(controls),
"iq": np.concatenate([treated, controls])
})
d.groupby("group").mean()
| iq | |
|---|---|
| group | |
| control | 100.357143 |
| treatment | 101.914894 |
Equal variances t-test#
from scipy.stats import ttest_ind
res_eqvar = ttest_ind(treated, controls, equal_var=True)
res_eqvar.statistic, res_eqvar.pvalue
(1.5586953301521096, 0.12269895509665575)
ci_eqvar = res_eqvar.confidence_interval(confidence_level=0.95)
[ci_eqvar.low, ci_eqvar.high]
[-0.42865302979133335, 3.5441545495481668]
Unequal variances t-test#
res_uneqvar = ttest_ind(treated, controls, equal_var=False)
res_uneqvar.statistic, res_uneqvar.pvalue
(1.622190457290228, 0.10975381983712831)
ci_uneqvar = res_uneqvar.confidence_interval(confidence_level=0.95)
[ci_uneqvar.low, ci_uneqvar.high]
[-0.3611847716497789, 3.476686291406612]
Linear model#
import statsmodels.formula.api as smf
res_ols = smf.ols("iq ~ 1 + C(group)", data=d).fit()
print(res_ols.summary())
OLS Regression Results
==============================================================================
Dep. Variable: iq R-squared: 0.027
Model: OLS Adj. R-squared: 0.016
Method: Least Squares F-statistic: 2.430
Date: Fri, 15 Nov 2024 Prob (F-statistic): 0.123
Time: 04:27:22 Log-Likelihood: -263.13
No. Observations: 89 AIC: 530.3
Df Residuals: 87 BIC: 535.2
Df Model: 1
Covariance Type: nonrobust
=========================================================================================
coef std err t P>|t| [0.025 0.975]
-----------------------------------------------------------------------------------------
Intercept 100.3571 0.726 138.184 0.000 98.914 101.801
C(group)[T.treatment] 1.5578 0.999 1.559 0.123 -0.429 3.544
==============================================================================
Omnibus: 46.068 Durbin-Watson: 2.025
Prob(Omnibus): 0.000 Jarque-Bera (JB): 553.494
Skew: 1.108 Prob(JB): 6.46e-121
Kurtosis: 15.014 Cond. No. 2.69
==============================================================================
Notes:
[1] Standard Errors assume that the covariance matrix of the errors is correctly specified.
Alternative linear model#
Using generalized least squares to reproduce the unequal variance case.
n_t, var_t = len(treated), treated.var()
n_c, var_c = len(controls), controls.var()
sigma2s = [var_t]*n_t + [var_c]*n_c
res_gls = smf.gls("iq ~ 1 + C(group)", data=d, sigma=sigma2s).fit()
print(res_gls.summary())
GLS Regression Results
==============================================================================
Dep. Variable: iq R-squared: 0.029
Model: GLS Adj. R-squared: 0.018
Method: Least Squares F-statistic: 2.629
Date: Fri, 15 Nov 2024 Prob (F-statistic): 0.109
Time: 04:27:22 Log-Likelihood: -248.41
No. Observations: 89 AIC: 500.8
Df Residuals: 87 BIC: 505.8
Df Model: 1
Covariance Type: nonrobust
=========================================================================================
coef std err t P>|t| [0.025 0.975]
-----------------------------------------------------------------------------------------
Intercept 100.3571 0.388 258.627 0.000 99.586 101.128
C(group)[T.treatment] 1.5578 0.961 1.622 0.109 -0.352 3.467
==============================================================================
Omnibus: 33.582 Durbin-Watson: 2.234
Prob(Omnibus): 0.000 Jarque-Bera (JB): 346.198
Skew: -0.648 Prob(JB): 6.67e-76
Kurtosis: 12.575 Cond. No. 2.79
==============================================================================
Notes:
[1] Standard Errors assume that the covariance matrix of the errors is correctly specified.
Bayesian equal variances model#
import bambi as bmb
mod_eqvar = bmb.Model("iq ~ 1 + group", data=d)
print(mod_eqvar)
Formula: iq ~ 1 + group
Family: gaussian
Link: mu = identity
Observations: 89
Priors:
target = mu
Common-level effects
Intercept ~ Normal(mu: 101.1798, sigma: 17.17)
group ~ Normal(mu: 0.0, sigma: 23.6275)
Auxiliary parameters
sigma ~ HalfStudentT(nu: 4.0, sigma: 4.718)
idata_eqvar = mod_eqvar.fit(draws=2000)
Auto-assigning NUTS sampler...
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [sigma, Intercept, group]
Sampling 2 chains for 1_000 tune and 2_000 draw iterations (2_000 + 4_000 draws total) took 3 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
import arviz as az
az.summary(idata_eqvar, kind="stats")
| mean | sd | hdi_3% | hdi_97% | |
|---|---|---|---|---|
| sigma | 4.751 | 0.370 | 4.056 | 5.433 |
| Intercept | 100.361 | 0.727 | 98.964 | 101.728 |
| group[treatment] | 1.555 | 0.986 | -0.424 | 3.333 |
Bayesian unequal variances model#
formula = bmb.Formula("iq ~ 1 + group", "sigma ~ group")
mod_uneqvar = bmb.Model(formula, data=d)
print(mod_uneqvar)
Formula: iq ~ 1 + group
sigma ~ group
Family: gaussian
Link: mu = identity
sigma = log
Observations: 89
Priors:
target = mu
Common-level effects
Intercept ~ Normal(mu: 101.1798, sigma: 17.17)
group ~ Normal(mu: 0.0, sigma: 23.6275)
target = sigma
Common-level effects
sigma_Intercept ~ Normal(mu: 0.0, sigma: 1.0)
sigma_group ~ Normal(mu: 0.0, sigma: 1.0)
idata_uneqvar = mod_uneqvar.fit(draws=2000)
Auto-assigning NUTS sampler...
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [Intercept, group, sigma_Intercept, sigma_group]
Sampling 2 chains for 1_000 tune and 2_000 draw iterations (2_000 + 4_000 draws total) took 5 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
az.summary(idata_uneqvar, kind="stats")
| mean | sd | hdi_3% | hdi_97% | |
|---|---|---|---|---|
| Intercept | 100.358 | 0.409 | 99.597 | 101.112 |
| group[treatment] | 1.585 | 0.974 | -0.259 | 3.422 |
| sigma_Intercept | 0.939 | 0.111 | 0.727 | 1.146 |
| sigma_group[treatment] | 0.851 | 0.152 | 0.579 | 1.141 |
Robust Bayesian Estimation#
formula = bmb.Formula("iq ~ 1 + group", "sigma ~ group")
mod_robust = bmb.Model(formula, family="t", data=d)
print(mod_robust)
Formula: iq ~ 1 + group
sigma ~ group
Family: t
Link: mu = identity
sigma = log
Observations: 89
Priors:
target = mu
Common-level effects
Intercept ~ Normal(mu: 101.1798, sigma: 17.17)
group ~ Normal(mu: 0.0, sigma: 23.6275)
Auxiliary parameters
nu ~ Gamma(alpha: 2.0, beta: 0.1)
target = sigma
Common-level effects
sigma_Intercept ~ Normal(mu: 0.0, sigma: 1.0)
sigma_group ~ Normal(mu: 0.0, sigma: 1.0)
idata_robust = mod_robust.fit(draws=1000)
az.summary(idata_robust, kind="stats")
Auto-assigning NUTS sampler...
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [nu, Intercept, group, sigma_Intercept, sigma_group]
Sampling 2 chains for 1_000 tune and 1_000 draw iterations (2_000 + 2_000 draws total) took 4 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
| mean | sd | hdi_3% | hdi_97% | |
|---|---|---|---|---|
| nu | 1.827 | 0.472 | 1.059 | 2.717 |
| Intercept | 100.527 | 0.207 | 100.158 | 100.928 |
| group[treatment] | 0.999 | 0.417 | 0.232 | 1.793 |
| sigma_Intercept | 0.008 | 0.197 | -0.353 | 0.385 |
| sigma_group[treatment] | 0.634 | 0.252 | 0.193 | 1.122 |